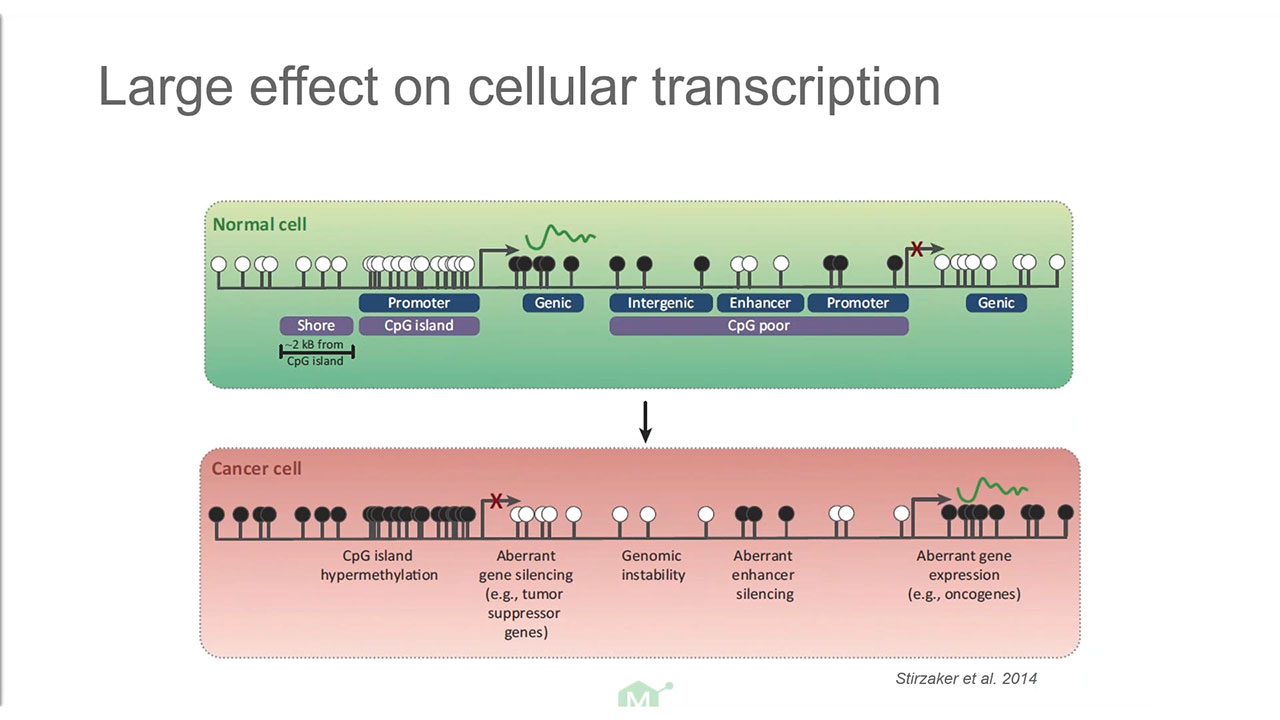
Slide-seq: A scalable technology for measuring genome-wide expression at high spatial resolution | Science

Fast-Seq, a universal method for rapid and inexpensive genomic validation of rAAV vectors in preclinical settings

TARGET-Seq: A Protocol for High-Sensitivity Single-Cell Mutational Analysis and Parallel RNA Sequencing - ScienceDirect

Div-Seq – A single nucleus RNA-Seq method reveals dynamics of rare adult newborn neurons in the CNS | RNA-Seq Blog

Dissecting Immune Circuits by Linking CRISPR-Pooled Screens with Single-Cell RNA-Seq - ScienceDirect

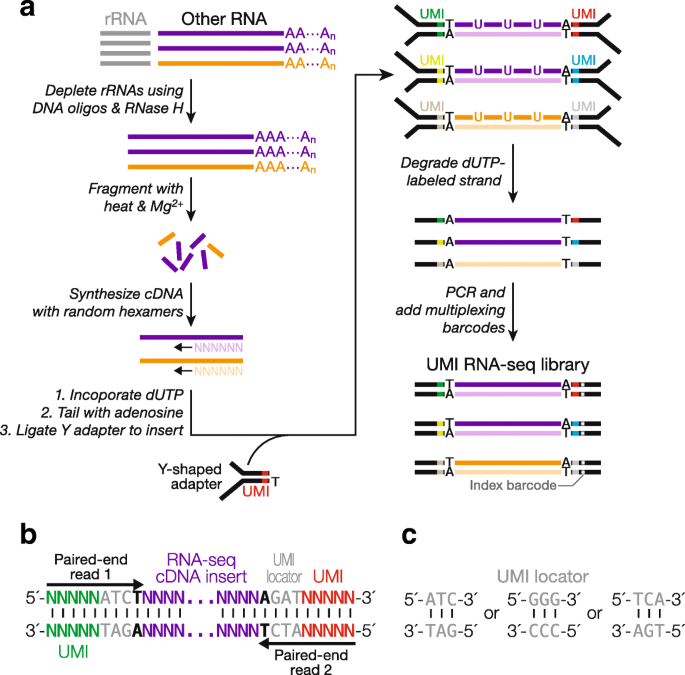
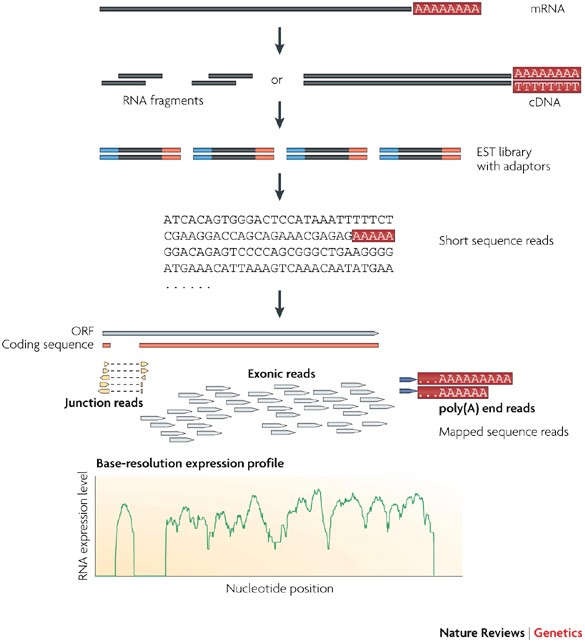
Overview of RNA-seq.A RNA fraction of interest is selected, fragmented... | Download Scientific Diagram

Introducing MAS-ISO-seq: A new long-read sequencing protocol for dramatically increasing RNA transcript isoform yield - Terra.Bio
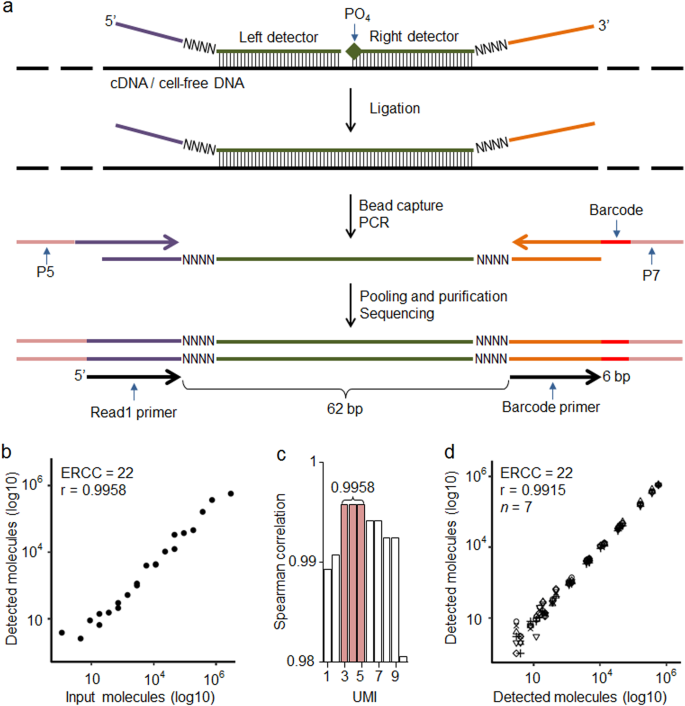
TAC-seq: targeted DNA and RNA sequencing for precise biomarker molecule counting | npj Genomic Medicine

DiMeLo-seq: a long-read, single-molecule method for mapping protein–DNA interactions genome wide | Nature Methods

DNase-seq: A High-Resolution Technique for Mapping Active Gene Regulatory Elements across the Genome from Mammalian Cells

A work flow for 3'end-seq and PacBio Iso-seq. (A) Multiple versions of... | Download Scientific Diagram

Introducing MAS-ISO-seq: A new long-read sequencing protocol for dramatically increasing RNA transcript isoform yield - Terra.Bio

Schematic diagram representing EXPRSS Tag-seq. (A) Library preparation... | Download Scientific Diagram
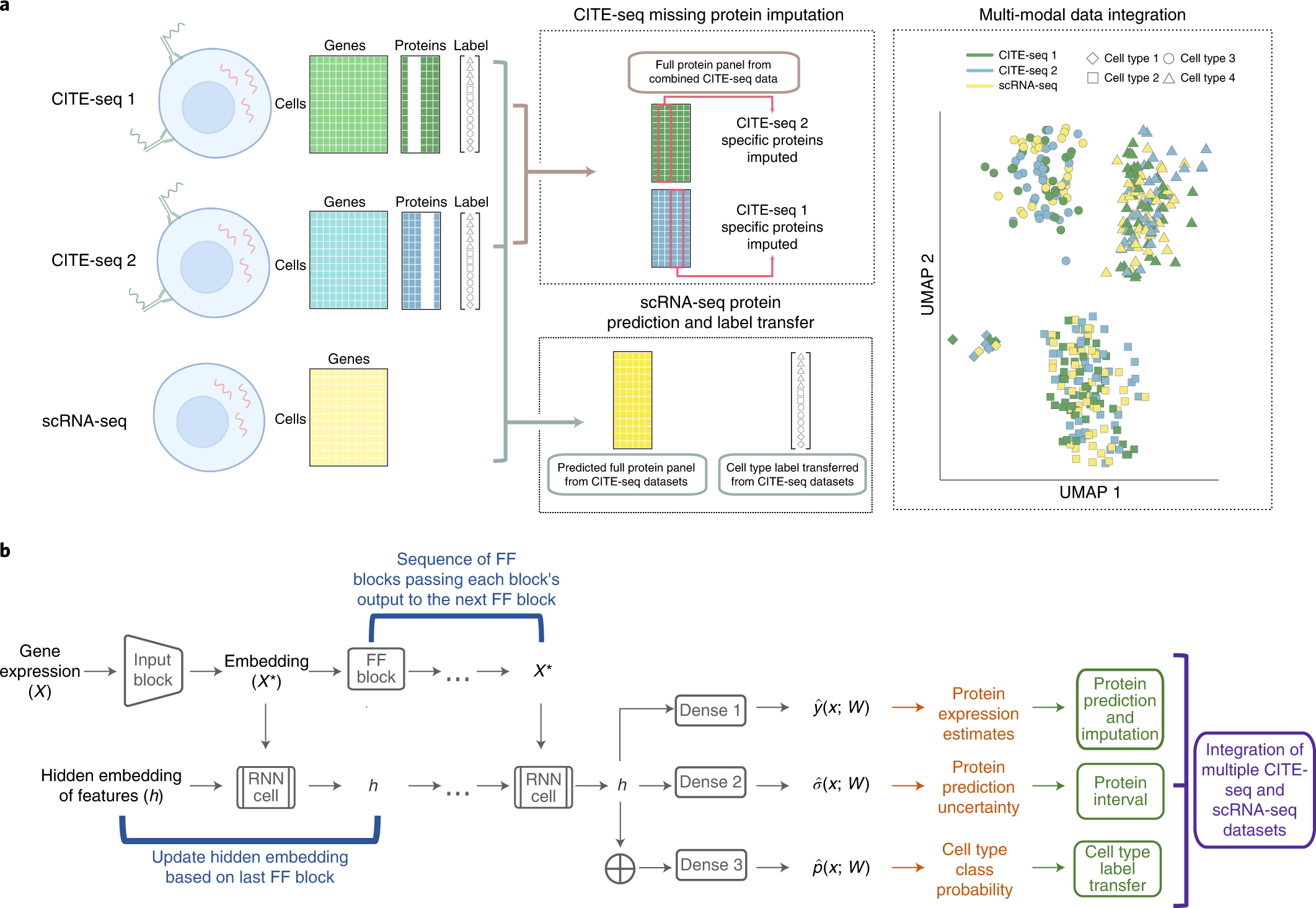
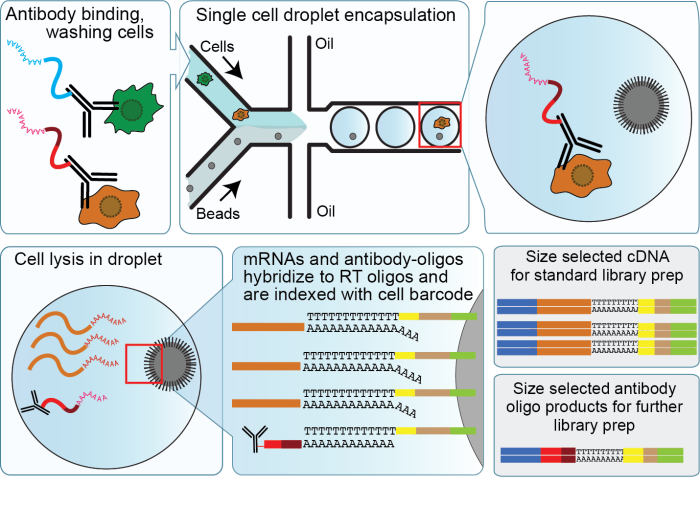
A multi-use deep learning method for CITE-seq and single-cell RNA-seq data integration with cell surface protein prediction and imputation | Nature Machine Intelligence